 When finished press the save button.I'm looking through a dataset set of images in ImageJ (a stack of. In this case you repeat the selection for all cells in the same images. If you select is for batch, you must use the rois button to add the transcription-site, the nucleus and a control region. It will contain the cropped cell and the selections T, N and C in the file RoiSet.zip. For each cell a folder with the name of the input image followed by _cn, where n is the number of the cell, will be created. When all cells in the image have been treated, continue with the next image. You can now repeat the procedure for the next cell. The cell window and the projection windows will be closed. You can transfer them between the images using Edit>Selection>Restore Selection (ctrl+shift+e). Note that you can use any projection to create the rois. Finally select a control region within the nucleus and add it in the same way. This time a roi with the label "N" is added. Than select the nucleus and press the +roi-button again. A roi with the label "T" is added to the roi-manager. Select the transcription-site and press the +roi-button. The cropped image of the cell will be saved and the projections will be displayed. When this is finished, switch back to the "MS2Quant_PP" toolset. Also note that you should not use the E-button (export cells) on this toolset. Note that only one region (the first) for each image will be taken into account. The next step is to use the Crop_4D_Cells toolset do define background regions. The process is displayed in a log-window: Run the concatenate stacks batch by clicking on the cs-button. is for batch: if selected the pre processing is done for MS2Quant batch processing, otherwise for manual processingįor a new experiment first set the number of z-sclices and time-frames in the options dialog.mark: a mark that allows to parse the filenames (the mark is "PST" in the example images).To become saturated in the contrast enhancement. This parameter fixes the number of pixels that are allowed enhance contrast (%saturated): when the projections are displayed, the display-contrast is automatically enhanced.time-frames: the number of time frames in the stack.z-slices: the number of z-slices in the stack.Open the options dialog by right-clicking on the ms2 button. The save-button saves the projection image and the rois for the batch-processing.The rois-button allows to add the selections for the transcription site, the nucleus and the control region for batch processing.The +roi-button allows to add the selections for the transcription site, the nucleus and the control region. 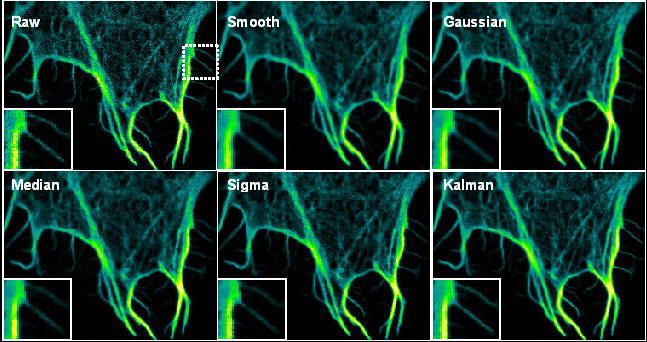
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |


 RSS Feed
RSS Feed
